_48.png)
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | Set_04_transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB.phenotype_transDMproB_versus_DMpreB.cls #transDMproB_versus_DMpreB_repos |
| Phenotype | phenotype_transDMproB_versus_DMpreB.cls#transDMproB_versus_DMpreB_repos |
| Upregulated in class | transDMproB |
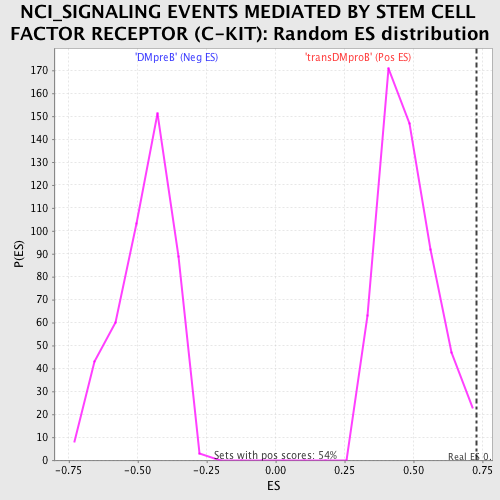
| GeneSet | NCI_SIGNALING EVENTS MEDIATED BY STEM CELL FACTOR RECEPTOR (C-KIT) |
| Enrichment Score (ES) | 0.72770506 |
| Normalized Enrichment Score (NES) | 1.5167575 |
| Nominal p-value | 0.009208103 |
| FDR q-value | 0.20227808 |
| FWER p-Value | 0.96 |
_48.png)
| PROBE | DESCRIPTION (from dataset) | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|---|
| 1 | KIT | 16823 | 40 | 3.755 | 0.1726 | Yes | ||
| 2 | GAB1 | 18828 | 64 | 3.185 | 0.3196 | Yes | ||
| 3 | STAT3 | 5525 9906 | 396 | 1.605 | 0.3764 | Yes | ||
| 4 | SH2B3 | 5001 16383 | 449 | 1.532 | 0.4449 | Yes | ||
| 5 | STAT1 | 3936 5524 | 531 | 1.404 | 0.5059 | Yes | ||
| 6 | CREBBP | 22682 8783 | 740 | 1.067 | 0.5443 | Yes | ||
| 7 | VAV1 | 23173 | 746 | 1.053 | 0.5930 | Yes | ||
| 8 | PDPK1 | 23097 | 914 | 0.896 | 0.6258 | Yes | ||
| 9 | PTEN | 5305 | 1062 | 0.791 | 0.6547 | Yes | ||
| 10 | STAT5A | 20664 | 1207 | 0.676 | 0.6784 | Yes | ||
| 11 | TEC | 16514 | 1558 | 0.496 | 0.6826 | Yes | ||
| 12 | CRKL | 4560 | 1800 | 0.391 | 0.6878 | Yes | ||
| 13 | GSK3B | 22761 | 1874 | 0.364 | 0.7009 | Yes | ||
| 14 | GRB2 | 20149 | 1927 | 0.347 | 0.7142 | Yes | ||
| 15 | BAD | 24000 | 1967 | 0.335 | 0.7277 | Yes | ||
| 16 | MAPK3 | 6458 11170 | 2440 | 0.214 | 0.7122 | No | ||
| 17 | RAF1 | 17035 | 3038 | 0.101 | 0.6848 | No | ||
| 18 | MAP2K1 | 19082 | 3174 | 0.087 | 0.6816 | No | ||
| 19 | RPS6KB1 | 7815 1207 13040 | 4229 | 0.026 | 0.6260 | No | ||
| 20 | PTPRO | 1093 17256 | 4323 | 0.023 | 0.6221 | No | ||
| 21 | EPOR | 19204 | 4385 | 0.022 | 0.6198 | No | ||
| 22 | MATK | 3425 19930 | 4494 | 0.019 | 0.6149 | No | ||
| 23 | SOS1 | 5476 | 4722 | 0.015 | 0.6034 | No | ||
| 24 | KITLG | 19889 3342 | 4811 | 0.014 | 0.5993 | No | ||
| 25 | PIK3R1 | 3170 | 4940 | 0.013 | 0.5930 | No | ||
| 26 | SNAI2 | 22848 | 5075 | 0.011 | 0.5863 | No | ||
| 27 | EPO | 8911 | 5254 | 0.010 | 0.5772 | No | ||
| 28 | MITF | 17349 | 5449 | 0.008 | 0.5671 | No | ||
| 29 | GRAP2 | 5113 9398 | 5964 | 0.006 | 0.5397 | No | ||
| 30 | DOK1 | 17104 1018 1177 | 5995 | 0.006 | 0.5384 | No | ||
| 31 | SPRED2 | 8539 4332 | 6754 | 0.003 | 0.4977 | No | ||
| 32 | FOXO3 | 19782 3402 | 6843 | 0.003 | 0.4931 | No | ||
| 33 | LYN | 16281 | 7120 | 0.002 | 0.4783 | No | ||
| 34 | PIK3CA | 9562 | 7760 | 0.001 | 0.4439 | No | ||
| 35 | BCL2 | 8651 3928 13864 4435 981 4062 13863 4027 | 9115 | -0.003 | 0.3711 | No | ||
| 36 | SHC1 | 9813 9812 5430 | 9137 | -0.003 | 0.3701 | No | ||
| 37 | PTPN11 | 5326 16391 9660 | 9713 | -0.004 | 0.3394 | No | ||
| 38 | GRB10 | 4799 | 11584 | -0.010 | 0.2391 | No | ||
| 39 | JAK2 | 23893 9197 3706 | 11807 | -0.012 | 0.2277 | No | ||
| 40 | AKT1 | 8568 | 13176 | -0.023 | 0.1551 | No | ||
| 41 | CBL | 19154 | 13815 | -0.033 | 0.1223 | No | ||
| 42 | SPRED1 | 4331 | 15362 | -0.105 | 0.0439 | No | ||
| 43 | MAP2K2 | 19933 | 15726 | -0.151 | 0.0314 | No | ||
| 44 | MAPK8 | 6459 | 16226 | -0.245 | 0.0159 | No | ||
| 45 | HRAS | 4868 | 16569 | -0.329 | 0.0128 | No | ||
| 46 | SOCS1 | 4522 | 16690 | -0.364 | 0.0232 | No | ||
| 47 | PTPN6 | 17002 | 17395 | -0.639 | 0.0150 | No | ||
| 48 | MAP4K1 | 18313 | 18009 | -1.088 | 0.0327 | No |
_49.png)